Welcome to Enhlink’s documentation!
Introduction
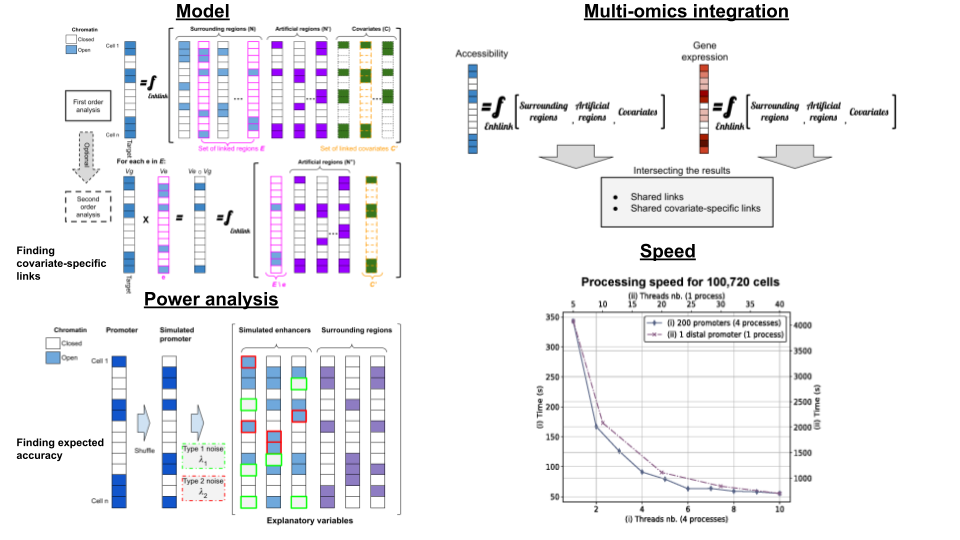
Enhlink is a fast, accurate, and efficient algorithm designed to perform linkage analysis, such as inferring enhanceer-promoter links, from single-cell ATAC-seq (scATAC-seq) datasets. It is also applicable to other single-cell genomic datasets (e.g., methylation, DNase, MNase, ChIP-seq) containing high-dimensional, sparse data with binary features.
Key Features
Versatile Application: Works with various single-cell genomic datasets, including scATAC-seq, methylation, DNase, MNase, and ChIP-seq.
Covariate Modeling: Allows modeling of covariates to enhance inference accuracy and can integrate into existing workflows.
Multi-Omic Integration: Optionally integrates multi-omic datasets, like gene expression, to boost the quality of linkage inference.
Predictive Feature Modeling: Models enhancers and covariates as predictive features of target accessibility or gene expression.
Modified Information Gain Score: Employs this metric within a random forest framework to estimate significant features linked to specific targets.
Second-Order Analysis: Identifies linkages that are specific to particular covariates.
Distal Enhancer-Gene Linkage: Capable of inferring linkages beyond proximal enhancers.
Simulation Workflow: Integrates a simulation workflow based on experimentally validated enhancer-promoter signals, improving prediction accuracy.
Introduction and Tutorials:
- Introduction and installation
- Tutorial 1: Simple processing of snATAC-seq data
- Tutorial 2: Use of simulated variables to estimate expected accuracy
- Tutorial 3: Create sparse matrices compatible with Enhlink
- Tutorial 4: Processing multi-omics data with Enhlink
- Tutorial 5: Enhlink advanced functionalities
- Inferring positively/negatively correlated linkage only
- Using an external feature matrix with Enhlink.
- Regularization and adjusting sensitivity / specificity
- Distal interactions inference
- using Enhgrid to distribute computation, test the robustness of results, and improve speed
- Filter links based on TAD domains
- Tutorial 6: Generating simulated matrices for method comparison
- Tutorial 7: Compatibility with other single-cell libraries ( Seurat, Signac, and SnapATAC)
- License
Access
The package is accessible at this link: https://gitlab.com/grouumf/enhlinktools/
Citation
Enhlink is published in Genome Biology: https://genomebiology.biomedcentral.com/articles/10.1186/s13059-024-03374-9
License
GNU GENERAL PUBLIC LICENSE Version 3, 29 June 2007
Support
If you are having issues, please let us know. You can reach us using the following email address:
Olivier Poirion, Ph.D. o.poirion@gmail.com (personal) or olivier.poirion@jax.org (current work)
Data availability
The PCHi-C data used in this manuscript are available with currently a limited access through GEO with the following accession: GSE214107. The data are planned to be released to the public the 31 December 2023. In the meantime, they will be available on demand. The Linkage atlases for the islet and adipose datasets are available as Figshare datasets fith the following DOIs: https://doi.org/10.6084/m9.figshare.22336033.v1 (adipose) and https://doi.org/10.6084/m9.figshare.22335919.v1 (islet).
Module API:
- enhgrid
- Library that compiles the enhgrid executable
- Index
- Variables
- func analyseOneGeneList
- func getGeneBucketsFromGene
- func getGeneBucketsFromPromoter
- func launchOneIterThread
- func main
- func mergeBucketResultsFile
- func mergeOneSetOfBucketFiles
- func processNGeneLists
- func reduce
- func splitGenesToBucket
- func stringToFloatArray
- func stringToIntArray
- func stringToMaxFeatTypeArray
- func testIfRequiredFilesExist
- type paramArrays
- enhlink
- enhlinkobject
- enhtools
- Library that compiles the enhtools executable
- Index
- Variables
- func checkIfLineCanBeSplitIntoPeaks
- func filterBedpe
- func filterWithBed
- func getPosFromOption
- func incompleteIntervalIntersect
- func intersectBedpeWithBedFile
- func intersectBedpes
- func intersectBedpes2
- func intersectBedpes3
- func intersectBedpes4
- func intersectOneInput
- func intersectWithBed
- func intervalIntersect
- func isIntersecting
- func loadPeakFile
- func main
- func writeHeader
- func writeOneInterToBuffer
- func writeStats
- type geneStats
- type geneStatsMap
- type intervalResults
- type matching
- type mergeFunc
- type metaMap
- type peakMeta
- type peakPair
- type scoreMerging
- type twoKeysBoolMap
- matrix
- Index
- Constants
- Variables
- func GetRandomBootstrapIndex
- func LoadIndexFileToIndex
- func LoadPeakDictsToIndex
- func Mean
- func Std
- func TestStringToPeak
- func TtestPval
- func diffMap
- func maxUintMap
- func minIndexSliceInt
- func minInt
- func processMtxHeader
- func reverseIndex
- func reverseIndexC
- type Attributes
- type Format
- type MatColFloatHash
- type MatColHash
- type SparseBoolMatrix
- type SparseFloatMatrix